Search
About Oryza glaberrima
Oryza glaberrima (African rice) is a cultivated grain distinct from its better known cousin Oryza sativa (Asian rice). African rice was independently domesticated ~3000 years ago in the Niger River Delta from its still extant progenitor, Oryza barthii. While lacking many of the agronomic and quality traits found in Asian rice, O. glaberrima is significant for its resistance to many pests and diseases and for its tolerance of drought and infertile soils. Interspecific crosses between African and Asian rice have produced cultivars with improved yield and quality traits, that have been adopted by many African countries to meet the growing need for rice as a staple food. From a scientific perspective the genome of O. glaberrima provides insight into the genetic basis of domestication and other traits by finding commonalities and differences with O. sativa. Similar to Asian rice, African rice is a diploid A-type genome, having 12 chromosomes and an estimated size of ~358 Mb.
Taxonomy ID 4538
Data source Oryza Genome Evolution Project
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
 Download alignments (EMF)
Download alignments (EMF)
Variation
What can I find? Short sequence variants.
 More about variation in Ensembl Plants
More about variation in Ensembl Plants
Regulation
What can I find? Microarray annotations.
 More about the Ensembl Plants microarray annotation strategy
More about the Ensembl Plants microarray annotation strategy
Gramene/Ensembl Genomes Annotation
Additional annotations generated by the Gramene and Ensembl Plants project include:
- Gene phylogenetic trees with other Gramene species.
- LastZ Whole Genome Alignment to Arabidopsis thaliana, Oryza sativa Japonica (IRGSP v1) and other Oryza AA genomes.
- Orthologue based DAGchainer synteny detection against other AA genomes.
- Mapping to the genome of multiple sequence-based feature sets using Gramene BLAT pipeline.
- Identification of various repeat features by programs such as RepeatMasker with MIPS and AGI repeat libraries, and Dust, TRF.
- Variation effect prediction with sequence ontology.




 Convert your data to Oryza_glaberrima_V1 coordinates
Convert your data to Oryza_glaberrima_V1 coordinates Display your data in Ensembl Plants
Display your data in Ensembl Plants

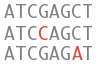

![Follow us on Twitter! [twitter logo]](/i/twitter.png)
